Detection of plasma HOXA7 methylation in the diagnosis of non-small cell lung cancer
-
摘要:
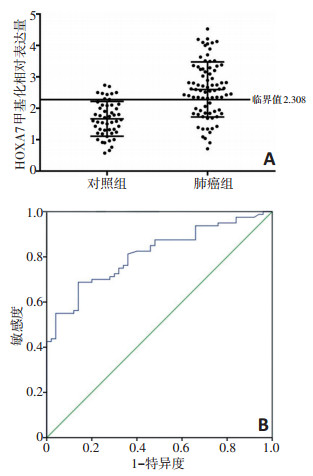
目的 检测非小细胞肺癌(non-small cell lung cancer, NSCLC)血浆中同源盒基因(homeobox, HOX)家族成员HOXA7基因启动子甲基化水平, 并评估HOXA7甲基化与肿瘤标志物血清癌胚抗原(carcinoembryonic antigen, CEA)、糖类抗原(carbohydrate antigen 199, CA199)和细胞角蛋白19片段抗原21-1(cytokeratin 19 fragment antigen 21-1, Cyfra21-1)水平辅助诊断NSCLC的价值。 方法 选取河南省肿瘤医院2012年1月至2016年12月住院的经病理组织学确诊为NSCLC患者80例为试验组, 50例健康人为对照组。采用定量甲基化特异性PCR(quantitative methylation-specific PCR, qMSP)检测血浆HOXA7甲基化水平, 电化学发光检测血浆CEA、CA199和Cyfra21-1水平。构建受试者工作特征曲线(ROC)分析各指标鉴别NSCLC的临床价值, 并分析HOXA7甲基化与临床病例参数、肿瘤标志物之间的关系。 结果 与健康对照组相比, NSCLC患者血浆中HOXA7基因甲基化水平异常增高(χ2=36.972, P < 0.0001), HOXA7甲基化诊断NSCLC的敏感度为68.8%(55/80), 特异度为86.0%(43/50)。血浆HOXA7甲基化与性别、年龄、有无吸烟史和病理类型无关(均P>0.05), 但与TNM分期有关(P < 0.05), 其中Ⅳ期患者血浆HOXA7甲基化水平最高。肿瘤标志物中Cyfra21-1诊断NSCLC的价值最高, 敏感度为70.00%, 特异度为90.00%。HOXA7甲基化联合三指标诊断NSCLC的效能最高(AUC=0.893, 敏感度73.75%, 特异度94%)。HOXA7甲基化与Cyfra21-1的相关系数最高(r=0.564, P < 0.05)。 结论 HOXA7基因甲基化水平对NSCLC的诊断有一定价值, 与NSCLC的恶性程度相关, 联合肿瘤标志物检测可以提高NSCLC的诊断效能。 -
关键词:
- 血浆 /
- HOXA7基因甲基化 /
- 非小细胞肺癌 /
- 肿瘤标志物
Abstract:Objective To detect the methylation levels of homeobox protein Hox A7 (HOXA7) gene promoter in the plasma of non-small cell lung cancer (NSCLC) and to study the value of HOXA7 methylation and serum levels of the tumor markers carcinoembryonic antigen (CEA), carbohydrate antigen 199 (CA199), and cytokeratin 19 fragment (Cyfra21-1) in the auxiliary diagnosis of NSCLC. Methods Plasma samples were collected from 80 patients with NSCLC and 50 healthy controls, which were enrolled in Henan Cancer Hospital from January 2012 to December 2016. The plasma HOXA7 methylation levels were detected by real-time quantitative methylation-specific PCR, and the plasma levels of CEA, CA199, and Cyfra21-1 were detected by electrochemical luminescence. The ROC curve was constructed to analyze the clinical value of each index in the differential diagnosis of NSCLC, and the relationship between HOXA7 methylation, clinical parameters, and tumor markers was analyzed. Results The serum levels of HOXA7 methylation in NSCLC patients were significantly higher (χ2=36.972, P < 0.000 1) than the healthy control group. The sensitivity and specificity of HOXA7 methylation in the diagnosis of NSCLC were 68.8% (55/ 80) and 86.0% (43/50), respectively. Plasma HOXA7 methylation was not related to gender, age, smoking history, or pathological type (all P>0.05), but was related to TNM stage (P < 0.05). The plasma HOXA7 methylation level of stage Ⅳ patients (81.8%, 18/22) was the highest. Cyfra21-1 had the highest value in the diagnosis of NSCLC among tumor markers with a sensitivity of 70.00% and a specificity of 90.00%. The combination of HOXA7 methylation with all three tumor markers had the highest efficiency (AUC=0.893, sensitivity 73.75%, specificity 94%) in the diagnosis of NSCLC. The correlation coefficient between HOXA7 methylation and Cyfra21-1 was the highest (r=0.564, P < 0.05). Conclusions HOXA7 gene methylation is related to the degree of metastasis of NSCLC and proves as an efficient diagnostic marker for NSCLC when combined with tumor marker detection. -
Key words:
- plasma /
- HOXA7 gene methylation /
- non-small cell lung cancer (NSCLC) /
- tumor marker
-
表 1 肺癌患者和健康人群的基本资料 n(%)

表 2 引物和探针序列

表 3 血浆HOXA7甲基化与临床病理参数的关系分析

表 4 血浆中3种肿瘤标志物和HOXA7甲基化对NSCLC的诊断价值评估

-
[1] Lee MC, Kadota K, Buitrago D, et al. Implementing the new IASLC/ATS/ ERS classification of lung adenocarcinomas: results from international and Chinese cohorts[J]. J Thorac Dis, 2014, 6(S5):S568-580. http://cn.bing.com/academic/profile?id=065855c524fea3006f8516668fa16ac9&encoded=0&v=paper_preview&mkt=zh-cn [2] Kangwan N, Park JM, Kim EH, et al. Chemoquiescence for ideal cancer treatment and prevention: where are we now[J]? J Cancer Prev, 2014, 19(2):89-96. doi: 10.15430/JCP.2014.19.2.89 [3] Liloglou T, Bediaga NG, Brown BR, et al. Epigenetic biomarkers in lung cancer[J]. Cancer Lett, 2014, 342(2):200-212. doi: 10.1016/j.canlet.2012.04.018 [4] Vendetti FP, Rudin CM. Epigenetic therapy in non- small- cell lung cancer: targeting DNA methyltransferases and histone deacetylases[J]. Expert Opin Biol Ther, 2013, 13(9):1273-1285. doi: 10.1517/14712598.2013.819337 [5] Ponomaryova AA, Rykova EY, Cherdyntseva NV, et al. Potentialities of aberrantly methylated circulating DNA for diagnostics and posttreatment follow-up of lung cancer patients[J]. Lung Cancer, 2013, 81 (3):397-403. doi: 10.1016/j.lungcan.2013.05.016 [6] Rauch T, Wang Z, Zhang X, et al. Homeobox genemethylation in lung cancer studied by genome- wide analysiswitha microarray- based methylated CpG island recovery assay[J]. Proc Natl Acad Sci, 2007, 104 (13):5527-5532. doi: 10.1073/pnas.0701059104 [7] Son JW, Jeong KJ, Jean WS, et al. Genome-wide combination profiling of DNA copy number and methylation for deciphering biomarkers in non-small cell lung cancer patients[J]. Cancer Lett, 2011, 311(1):29-37. doi: 10.1016/j.canlet.2011.06.021 [8] Hammerman PS, Lawrence MS, Voet D, et al. Comprehensive genomic characterization of squamous cell lung cancers. Cancer Genome Atlas Research Network[J]. Nature, 2012, 489(7417):519-525. doi: 10.1038/nature11404 [9] Hulbert A, Jusue-Torres I, Stark A, Early detection of lung cancer using DNA promoter hypermethylation in plasma and sputum[J]. Clin Cancer Res, 2017, 23(8):1998-2005. doi: 10.1158/1078-0432.CCR-16-1371 [10] Szpechcinski A, Chorostowska-Wynimko J, Kupis W, et al. Quantitative analysis of free-circulating DNA in plasma of patients with resectable NSCLC[J]. Expert Opin Biol Ther, 2012, 12(S1):S3-9. http://cn.bing.com/academic/profile?id=0fa4f2af4769dfd405851c36835e5fe9&encoded=0&v=paper_preview&mkt=zh-cn [11] Ulivi P, Mercatali L, Casoni GL, et al. Multiple marker detection in peripheral blood for NSCLC diagnosis[J]. PLoS One, 2013, 8(2):e57401. http://d.old.wanfangdata.com.cn/OAPaper/oai_pubmedcentral.nih.gov_3582604 [12] Rezsohazy R, Saurin AJ, Maurel-Zaffran C, et al. Cellular and molecular insights into Hox protein action[J]. Development, 2015, 142(7): 1212- 1227. doi: 10.1242/dev.109785 [13] Bhatlekar S, Fields JZ, Boman BM. HOX genes and their role in the development of human cancers[J]. J Mol Med, 2014, 92(8):811-823. doi: 10.1007/s00109-014-1181-y [14] 毕琦玲, 周立红, 韩毅敏.HOX基因与妇科肿瘤的研究进展[J].实用肿瘤学杂志, 2018, 32(2):140-143. http://d.old.wanfangdata.com.cn/Periodical/syzlxzz201802014 [15] Diaz-Lagares A, Mendez-Gonzalez J, Hervas D, et al. A novel epigenetic signature for early diagnosis in lung cancer[J]. Clin Cancer Res, 2016, 22(13):3361-3371. doi: 10.1158/1078-0432.CCR-15-2346 [16] Ettinger DS, Akerley W, Borghaei H, et al. National comprehensive cancer network. Non-small cell lung cancer[J]. J Natl Compr Canc Netw, 2013, 11(6):645-653. doi: 10.6004/jnccn.2013.0084 [17] Kalemkerian GP, Akerley W, Bogner P, et al. National comprehensive cancer network. Small cell lung cancer[J]. J Natl Compr Canc Netw, 2013, 11(1):78-98. doi: 10.6004/jnccn.2013.0011 [18] Liu L, Teng J, Zhang L, et al. The combination of the tumor markers suggests the histological diagnosis of lung cancer[J]. Biomed Res Int, 2017, 2017:2013989. http://cn.bing.com/academic/profile?id=09fea7ba6e4a61667207d7b60c48aed1&encoded=0&v=paper_preview&mkt=zh-cn [19] Okamura K, Takayama K, Izumi M, et al. Diagnostic value of CEA and CYFRA 21-1 tumor markers in primary lung cancer[J]. Lung Cancer, 2013, 80(1):45-49. http://cn.bing.com/academic/profile?id=3b95b0ee3de1041012e36a5271ab0dbc&encoded=0&v=paper_preview&mkt=zh-cn [20] 冯香梅, 王国庆, 陈瑛, 等.血清肿瘤标志物在肺癌诊断中的应用价值[J].中国肿瘤临床, 2010, 37(6):331-334. doi: 10.3969/j.issn.1000-8179.2010.06.009 [21] Li X, Asmitananda T, Gao L, et al. Biomarkers in the lung cancer diagnosis: a clinical perspective[J]. Neoplasma, 2012, 59(5):500-507. doi: 10.4149/neo_2012_064 -



 下载:
下载: