Research progress on tumor protein biomarkers using high-throughput proteomics based on mass spectrometry
-
摘要: 蛋白质组学研究虽已开展20余年,早期研究基于Western blot技术,后来采用质谱技术鉴定蛋白质。但最初质谱技术的测序深度和鉴定蛋白质数量有限,使蛋白质组学研究遇到瓶颈。近年来,随着质谱技术的飞速发展,实现了高通量的蛋白质组学鉴定,从此蛋白质组学研究进入新时代。目前,高通量蛋白质组学技术在肿瘤领域的应用主要包括揭示肿瘤发生发展机制、寻找特异性生物标志物、阐明耐药性产生机制和发现新治疗靶点等。肿瘤的早期发现和诊断有助于及时地进行医疗干预,从而大幅提高患者的生存率及生存质量。国内外研究已发现了很多候选肿瘤生物标志物,但仅极少数应用于临床。因此,仅针对探索肿瘤特异性生物标志物,本文选取并分析高通量蛋白质组学技术应用较为广泛深入、且发病率/死亡率较高的肺癌、乳腺癌、结直肠癌和肝癌4种肿瘤类型。在不同肿瘤类型中筛选发表期刊影响因子较高、经扩大样本量验证或功能验证、证据较强的研究。介绍基于质谱的高通量蛋白质组学技术和常用标本类型,重点综述在上述4种高发癌症中,基于该技术发现的可能用于早期诊断、预测预后、靶向治疗的蛋白质标志物,旨在为实现肿瘤精准诊疗提供新的理论依据。Abstract: Proteomics research has been performed for more than 20 years. Early research was based on simple Western blot technology, and later, mass spectrometry was used to identify proteins. However, the limited sequencing depth and the number of identified proteins in initial mass spectrometry technology have caused bottlenecks in proteomics research. In recent years, with the rapid development of mass spectrometry technology, high-throughput proteomics identification has been achieved, and proteomics research has entered a new era. At present, the application of high-throughput proteomics technology in the field of cancer mainly includes revealing the mechanism of tumorigenesis and development, searching for specific biomarkers, elucidating the mechanism of drug resistance, and identifying new therapeutic targets. Early detection and diagnosis of tumors are helpful for timely medical intervention, greatly improving the survival rate and quality of life. Recently, several candidate tumor biomarkers have been identified, but only a few have been used clinically. Here, aiming to explore tumor-specific biomarkers, we selected four high morbidity/mortality rate tumors with extensive application of high-throughput proteomics technology, such as lung, breast, colorectal, and liver cancer. We screened recently published studies from journals with high impact factors, evaluated by their expanded sample size or functional verification, and strong evidence in different tumor types. This article first briefly introduces mass spectrometry-based high-throughput proteomics technology and commonly used specimen types, and then focuses on reviewing the protein markers that may be used for early diagnosis, prognosis, and targeted therapy in the above four high-incidence cancers, aiming to provide a new theoretical basis for accurate diagnosis and treatment of tumors.
-
Key words:
- proteomics /
- mass spectrometry /
- tumor /
- biomarkers
-
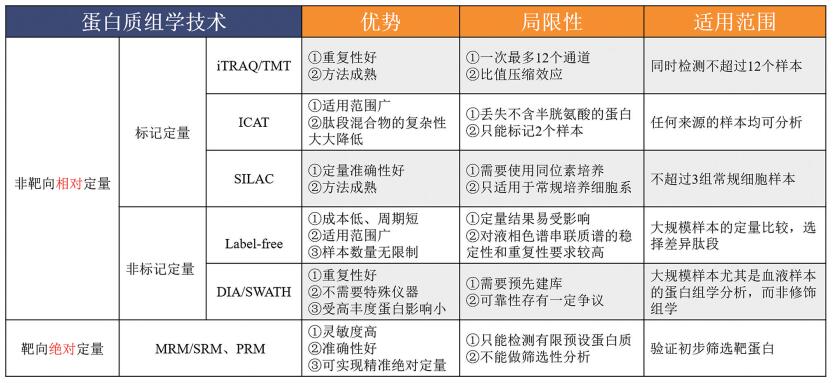
表 1 基于MS的高通量蛋白质组学技术发现不同肿瘤蛋白标志物

-
[1] Ferlay J, Soerjomataram I, Dikshit R, et al. Cancer incidence and mortality worldwide:sources, methods and major patterns in GLOBOCAN 2012[J]. Int J Cancer, 2015, 136(5):E359-E86. http://cn.bing.com/academic/profile?id=92998753a0e7f31e127db34e72d37cd5&encoded=0&v=paper_preview&mkt=zh-cn [2] Thun MJ, Delancey JO, Center MM, et al. The global burden of cancer:priorities for prevention[J]. Carcinogenesis, 2010, 31(1):100-110. http://cn.bing.com/academic/profile?id=3d9df2669d3bc9757afdb6e5ba0e3144&encoded=0&v=paper_preview&mkt=zh-cn [3] Ciocan-Cartita CA, Jurj A, Buse M, et al. The relevance of mass spectrometry analysis for personalized medicine through its successful application in cancer "Omics"[J]. Int J Mol Sci, 2019, 20(10):E2576. http://cn.bing.com/academic/profile?id=7b1e6a25dbbff08510b8bf26d9df7622&encoded=0&v=paper_preview&mkt=zh-cn [4] Belczacka I, Latosinska A, Metzger J, et al. Proteomics biomarkers for solid tumors:Current status and future prospects[J]. Mass Spectrom Rev, 2019, 38(1):49-78. http://cn.bing.com/academic/profile?id=7f946359a0b46433c4385f8206582520&encoded=0&v=paper_preview&mkt=zh-cn [5] Kowalczyk T, Ciborowski M, Kisluk J, et al. Mass spectrometry based proteomics and metabolomics in personalized oncology[J]. Biochim Biophys Acta Mol Basis Dis, 2020, 1866(5):165690. https://www.sciencedirect.com/science/article/pii/S0925443920300296 [6] Li N, Zhan X. Mass spectrometry-based mitochondrial proteomics in human ovarian cancers[J]. Mass Spectrom Rev, 2020[Epub ahead of print]. doi: 10.1002/mas.21618 [7] Calderon-Celis F, Encinar JR, Sanz-Medel A. Standardization approaches in absolute quantitative proteomics with mass spectrometry[J]. Mass Spectrom Rev, 2018, 37(6):715-737. http://www.wanfangdata.com.cn/details/detail.do?_type=perio&id=b13e9bb16c60ec8db3c2d19e82691051 [8] Neagu AN. Proteome imaging:from classic to modern mass spectrometry-based molecular histology[J]. Adv Exp Med Biol, 2019, 1140:55-98. http://cn.bing.com/academic/profile?id=9fc4ec6d59b48e415df5798e0700f17b&encoded=0&v=paper_preview&mkt=zh-cn [9] Siegel RL, Miller KD, Jemal A. Cancer statistics, 2019[J]. CA Cancer J Clin, 2019, 69(1):7-34. http://d.old.wanfangdata.com.cn/Periodical/zgazyj201905012 [10] Dai L, Qu Y, Li J, et al. Serological proteome analysis approach-based identification of ENO1 as a tumor-associated antigen and its autoantibody could enhance the sensitivity of CEA and CYFRA 21-1 in the detection of non-small cell lung cancer[J]. Oncotarget, 2017, 8(22):36664-36673. http://cn.bing.com/academic/profile?id=fc721480418f7830912b09afb2e11304&encoded=0&v=paper_preview&mkt=zh-cn [11] Jin Y, Yang Y, Su Y, et al. Identification a novel clinical biomarker in early diagnosis of human non-small cell lung cancer[J]. Glycoconj J, 2019, 36(1):57-68. http://www.wanfangdata.com.cn/details/detail.do?_type=perio&id=7ce8896c50b8927a3b53b03bc30eedc3 [12] Bohnenberger H, Kaderali L, Strobel P, et al. Comparative proteomics reveals a diagnostic signature for pulmonary head-and-neck cancer metastasis[J]. EMBO Mol Med, 2018, 10(9):e8428. http://cn.bing.com/academic/profile?id=12f14a90b873dd3f8a5c0e000a340d4a&encoded=0&v=paper_preview&mkt=zh-cn [13] Wang K, Li H, Chen R, et al. Combination of CALR and PDIA3 is a potential prognostic biomarker for non-small cell lung cancer[J]. Oncotarget, 2017, 8(57):96945-96957. http://cn.bing.com/academic/profile?id=ddd0840e74a68e63f8b8f62ba617595c&encoded=0&v=paper_preview&mkt=zh-cn [14] Bottger F, Schaaij-Visser TB, de Reus I, et al. Proteome analysis of nonsmall cell lung cancer cell line secretomes and patient sputum reveals biofluid biomarker candidates for cisplatin response prediction[J]. J Proteomics, 2019, 196:106-119. https://www.researchgate.net/publication/330746488_Proteome_analysis_of_non-small_cell_lung_cancer_cell_line_secretomes_and_patient_sputum_reveals_biofluid_biomarker_candidates_for_cisplatin_response_prediction [15] Sung HJ, Ahn JM, Yoon YH, et al. Quiescin sulfhydryl oxidase 1(qsox1) secreted by lung cancer cells promotes cancer metastasis[J]. Int J Mol Sci, 2018, 19(10):E3213. http://cn.bing.com/academic/profile?id=49a34aed62f0c2ccbddf457d7cabc03b&encoded=0&v=paper_preview&mkt=zh-cn [16] Parsons J, Francavilla C. 'Omics approaches to explore the breast cancer landscape[J]. Front Cell Dev Biol, 2019, 22(7):395. http://cn.bing.com/academic/profile?id=7f8a151937ae0319f83129cca4e3f13e&encoded=0&v=paper_preview&mkt=zh-cn [17] Tyanova S, Albrechtsen R, Kronqvist P, et al. Proteomic maps of breast cancer subtypes[J]. Nat Commun, 2016, 7:10259. http://cn.bing.com/academic/profile?id=2af02545463df5eecaee9039cf06fb7a&encoded=0&v=paper_preview&mkt=zh-cn [18] Valo I, Raro P, Boissard A, et al. OLFM4 expression in ductal carcinoma in situ and in invasive breast cancer cohorts by a SWATH-based proteomic approach[J]. Proteomics, 2019, 19(21-22):e1800446. http://cn.bing.com/academic/profile?id=800a79f6a359bb220cc262b58705cbb4&encoded=0&v=paper_preview&mkt=zh-cn [19] Pedersen MH, Hood BL, Ehmsen S, et al. CYPOR is a novel and independent prognostic biomarker of recurrence-free survival in triplenegative breast cancer patients[J]. Int J Cancer, 2019, 144(3):631-640. doi: 10.1002/ijc.31798 [20] Zeng L, Deng X, Zhong J, et al. Prognostic value of biomarkers EpCAM and alphaB-crystallin associated with lymphatic metastasis in breast cancer by iTRAQ analysis[J]. BMC Cancer, 2019, 19(1):831. https://www.researchgate.net/publication/335372655_Prognostic_value_of_biomarkers_EpCAM_and_aB-crystallin_associated_with_lymphatic_metastasis_in_breast_cancer_by_iTRAQ_analysis?origin=publication_list [21] Westbrook JA, Cairns DA, Peng J, et al. CAPG and GIPC1:breast cancer biomarkers for bone metastasis development and treatment[J]. J Natl Cancer Inst, 2016, 108:4. http://cn.bing.com/academic/profile?id=4a058859e4794f53f02457f8d6733743&encoded=0&v=paper_preview&mkt=zh-cn [22] Westbrook JA, Wood SL, Cairns DA, et al. Identification and validation of DOCK4 as a potential biomarker for risk of bone metastasis development in patients with early breast cancer[J]. J Pathol, 2019, 247(3):381-391. http://www.wanfangdata.com.cn/details/detail.do?_type=perio&id=10.1002/path.5197 [23] de Boer HR, Pool M, Joosten E, et al. Quantitative proteomics analysis identifies MUC1 as an effect sensor of EGFR inhibition[J]. Oncogene, 2019, 38(9):1477-1488. http://cn.bing.com/academic/profile?id=9ac12f6dcd69dff32062618cfa8a57e4&encoded=0&v=paper_preview&mkt=zh-cn [24] Pellegrino M, Rizza P, Dona A, et al. FoxO3a as a positive prognostic marker and a therapeutic target in tamoxifen-resistant breast cancer[J]. Cancers(Basel), 2019, 11:12. http://cn.bing.com/academic/profile?id=87ac1f7a62e506bef596443fc789b5df&encoded=0&v=paper_preview&mkt=zh-cn [25] Alves Martins BA, De Bulhoes GF, Cavalcanti IN, et al. Biomarkers in colorectal cancer:the role of translational proteomics research[J]. Front Oncol, 2019, 9:1284. https://www.researchgate.net/publication/337563524_Biomarkers_in_Colorectal_Cancer_The_Role_of_Translational_Proteomics_Research [26] Yu J, Zhai X, Li X, et al. Identification of MST1 as a potential early detection biomarker for colorectal cancer through a proteomic approach[J]. Sci Rep, 2017, 7(1):14265. http://cn.bing.com/academic/profile?id=de3423daf896e2c27feaebaff0d7a3a3&encoded=0&v=paper_preview&mkt=zh-cn [27] Duan B, Bai J, Qiu J, et al. Histone-lysine N-methyltransferase SETD7 is a potential serum biomarker for colorectal cancer patients[J]. EBioMedicine, 2018, 37:134-143. http://cn.bing.com/academic/profile?id=3177adf329017ed933302d762f4758b7&encoded=0&v=paper_preview&mkt=zh-cn [28] Bhardwaj M, Gies A, Weigl K, et al. Evaluation and validation of plasma proteins using two different protein detection methods for early detection of colorectal cancer[J]. Cancers (Basel), 2019, 25(11):10. http://cn.bing.com/academic/profile?id=a1a8f628caa5ea8866caadd9d915d5fa&encoded=0&v=paper_preview&mkt=zh-cn [29] Mori K, Toiyama Y, Otake K, et al. Successful identification of a predictive biomarker for lymph node metastasis in colorectal cancer using a proteomic approach[J]. Oncotarget, 2017, 8(63):106935-106947. http://cn.bing.com/academic/profile?id=09cfde80ce560e70af18debcc329e900&encoded=0&v=paper_preview&mkt=zh-cn [30] Mori K, Toiyama Y, Otake K, et al. Proteomics analysis of differential protein expression identifies heat shock protein 47 as a predictive marker for lymph node metastasis in patients with colorectal cancer[J]. Int J Cancer, 2017, 140(6):1425-1435. http://cn.bing.com/academic/profile?id=9938d04cfaff8951ad03274e8c26fa8f&encoded=0&v=paper_preview&mkt=zh-cn [31] Kirana C, Peng L, Miller R, et al. Combination of laser microdissection, 2D-DIGE and MALDI-TOF MS to identify protein biomarkers to predict colorectal cancer spread[J]. Clin Proteomics, 2019, 22(16):3. http://cn.bing.com/academic/profile?id=9cff4f62b4422f9be5587370ce1201ba&encoded=0&v=paper_preview&mkt=zh-cn [32] Guo J, Zhu C, Yang K, et al. Poly(C)-binding protein 1 mediates drug resistance in colorectal cancer[J]. Oncotarget, 2017, 8(8):13312-13319. http://cn.bing.com/academic/profile?id=3de289c21758364ac374b3c173d040cc&encoded=0&v=paper_preview&mkt=zh-cn [33] Lee YS, Ko E, Yoon EL, et al. Multiplexed proteomic approach for identification of serum biomarkers in hepatocellular carcinoma patients with normal AFP[J]. J Clin Med, 2020, 23(9):2. http://cn.bing.com/academic/profile?id=629286bd7b2318ab611aa8ef0fa2d3d8&encoded=0&v=paper_preview&mkt=zh-cn [34] Zhan Z, Guan Y, Mew K, et al. Urine alpha-fetoprotein and orosomucoid 1 as biomarkers of hepatitis B virus-associated hepatocellular carcinoma[J]. Am J Physiol Gastrointest Liver Physiol, 2020, 318(2):G305-G312. http://cn.bing.com/academic/profile?id=1e7de935391ff00035d3ca8ee611796d&encoded=0&v=paper_preview&mkt=zh-cn [35] Gao Q, Zhu H, Dong L, et al. Integrated proteogenomic characterization of hbv-related hepatocellular carcinoma[J]. Cell, 2019, 179(2):561-577. http://cn.bing.com/academic/profile?id=6baef17406879ece9ad5da4f3a3fba9a&encoded=0&v=paper_preview&mkt=zh-cn [36] Hindupur SK, Colombi M, Fuhs SR, et al. The protein histidine phosphatase LHPP is a tumour suppressor[J]. Nature, 2018, 555(7698):678-682. http://www.wanfangdata.com.cn/details/detail.do?_type=perio&id=2675fcddada03f8cf23116ac5977fcc4 [37] Jiang Y, Sun A, Zhao Y, et al. Proteomics identifies new therapeutic targets of early-stage hepatocellular carcinoma[J]. Nature, 2019, 567(7747):257-261. http://www.wanfangdata.com.cn/details/detail.do?_type=perio&id=a9f5eab176f0fd03a6d1d3a2ee42d038 [38] Kim H, Yu SJ, Yeo I, et al. prediction of response to sorafenib in hepatocellular carcinoma:a putative marker panel by multiple reaction monitoring-mass spectrometry(MRM-MS)[J]. Mol Cell Proteomics, 2017, 16(7):1312-1323. http://cn.bing.com/academic/profile?id=488c129afe70c05bcae0151a94dcb9b0&encoded=0&v=paper_preview&mkt=zh-cn [39] Le Large TYS, Meijer LL, Paleckyte R, et al. Combined expression of plasma thrombospondin-2 and CA19-9 for diagnosis of pancreatic cancer and distal cholangiocarcinoma:a proteome approach[J]. Oncologist, 2020, 25(4):e634-e643. http://cn.bing.com/academic/profile?id=b1c23ae402d50d7827dd530cc0d6aa28&encoded=0&v=paper_preview&mkt=zh-cn [40] Yoneyama T, Ohtsuki S, Honda K, et al. Identification of IGFBP2 and IGFBP3 as compensatory biomarkers for CA19-9 in early-stage pancreatic cancer using a combination of antibody-based and LC-MS/MSbased proteomics[J]. PLoS One, 2016, 11(8):e0161009. doi: 10.1371/journal.pone.0161009 [41] Qi Y, Zhang Y, Peng Z, et al. SERPINH1 overexpression in clear cell renal cell carcinoma:association with poor clinical outcome and its potential as a novel prognostic marker[J]. J Cell Mol Med, 2018, 22(2):1224-1235. http://cn.bing.com/academic/profile?id=05a04a62db7acaa010213857740ba6a4&encoded=0&v=paper_preview&mkt=zh-cn [42] Yang T, Xu P, Gu L, et al. Quantitative assessment of serum heat shock protein 27 for the diagnosis of epithelial ovarian cancer using targeted proteomics coupled with immunoaffinity enrichment[J]. Clin Chim Acta, 2019, 489:96-102. http://www.wanfangdata.com.cn/details/detail.do?_type=perio&id=c82c16bd16428f974bdf011a19209403 [43] Farrokhi Yekta R, Arefi Oskouie A, Rezaei Tavirani M, et al. Decreased apolipoprotein A4 and increased complement component 3 as potential markers for papillary thyroid carcinoma:A proteomic study[J]. Int J Biol Markers, 2018, 33(4):455-462. http://cn.bing.com/academic/profile?id=952af73b94cd01b904bbd1076f1d2519&encoded=0&v=paper_preview&mkt=zh-cn [44] Aleckovic M, Wei Y, Leroy G, et al. Identification of nidogen 1 as a lung metastasis protein through secretome analysis[J]. Genes Dev, 2017, 31(14):1439-1455. http://www.wanfangdata.com.cn/details/detail.do?_type=perio&id=64454a9c94afdcd8105ef2f79d657713 -



 下载:
下载: