-
摘要: 尤文肉瘤(ewing sarcoma,ES)和尤文样肉瘤均是侵袭性较强的小圆细胞肉瘤,两者的形态学和免疫表型相似,但发病部位、临床表现、预后及治疗方案却存在明显差异。尤文样肉瘤缺乏ES的典型分子特征,越来越多的证据表明,尤文样肉瘤是具有独特疾病特征的独立肉瘤,而不仅是ES的亚型。分子遗传学检测极大地促进了对小圆细胞肿瘤分类的重新评估,本文将就ES及尤文样肉瘤的主要病理分型—CIC重排肉瘤、BCOR重排肉瘤、EWSR1与非ETS家族基因重排的肉瘤和未分化小圆细胞肉瘤的最新研究进展进行综述。Abstract: Ewing's sarcoma and Ewing-like sarcoma are highly invasive sarcomas predominantly characterized by the presence of small round cells. Although certain morphological and immunophenotypical characteristics are similar between the two sarcomas, there are obvious differences in the disease sites, clinical manifestations, prognoses, and treatment responses. Additionally, Ewing-like sarcomas lack the typical molecular characteristics of Ewing's sarcoma. A growing body of evidence shows that Ewing-like sarcomas constitute a different disease type with unique characteristics, and are not just a subtype of Ewing's sarcoma. The discovery of novel molecular genetic tests has significantly aided the reevaluation of the classification of small round cell tumors. This article aims to review the recent progress in the research on Ewing's sarcoma and the major subclasses of Ewing-like sarcoma characterized thus far: CIC rearrangement sarcoma, BCOR rearrangement sarcoma, non-ETS family gene rearrangement sarcoma, and undifferentiated small round cell sarcoma.
-
Key words:
- Ewing's sarcoma /
- Ewing-like sarcoma /
- differential diagnosis
-
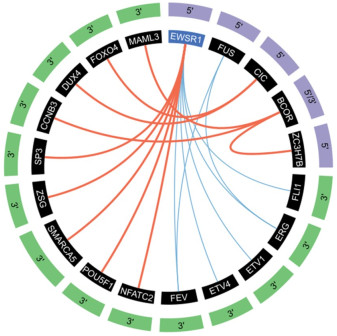
图 1 经典ES和尤文样肿瘤融合概况[10]
经典ES的基因融合以蓝色线条描绘,涉及RNA结合蛋白Tet家族(EWSR1和FUS)和ETS转录因子家族(最常见的是FLI1)之间的融合。尤文样肉瘤的融合以红色粗体表示。5'融合伙伴以紫色突出显示,3'融合伙伴以绿色突出显示
表 1 经典ES融合

表 2 尤文样肉瘤融合

表 3 ES与尤文样肉瘤的形态学、免疫表型及预后等异同汇总

-
[1] Marley EF, Liapis H, Humphrey PA, et al. Primitive neuroectodermal tumor of the kidney——another enigma:a pathologic, immunohistochemical, and molecular diagnostic study[J]. Am J Surg Pathol, 1997, 21(3):354-359. http://jcp.bmj.com/lookup/external-ref?access_num=9060607&link_type=MED&atom=%2Fjclinpath%2F58%2F10%2F1051.atom [2] Komforti MK, Sokolovskaya E, D'Agostino CA, et al. Extra-osseous Ewing sarcoma of the pancreas:case report with radiologic, pathologic, and molecular correlation, and brief review of the literature[J]. Virchows Arch, 2018, 473(3):361-369. http://cn.bing.com/academic/profile?id=ae4597cfaf177cf37ccda9eda9c43705&encoded=0&v=paper_preview&mkt=zh-cn [3] Dedeurwaerdere F, Giannini C, Sciot R, et al. Primary peripheral PNET/Ewing's sarcoma of the dura:a clinicopathologic entity distinct from central PNET[J]. Mod Pathol, 2002, 15(6):673-678. http://cn.bing.com/academic/profile?id=fcf0b785e2b7fea311a30822b6c1b574&encoded=0&v=paper_preview&mkt=zh-cn [4] Hasegawa SL, Davison JM, Rutten A, et al. Primary cutaneous Ewing's sarcoma:immunophenotypic and molecular cytogenetic evaluation of five cases[J]. Am J Surg Pathol, 1998, 22(3):310-318. http://europepmc.org/abstract/MED/9500772 [5] Charville GW, Wang WL, Ingram DR, et al. EWSR1 fusion proteins mediate PAX7 expression in Ewing sarcoma[J]. Mod Pathol, 2017, 30(9):1312-1320. http://www.wanfangdata.com.cn/details/detail.do?_type=perio&id=223733e9d334fc8367cf490a4dac039d [6] Yoshida A, Sekine S, Tsuta K, et al. NKX2.2 is a useful immunohistochemical marker for Ewing sarcoma[J]. Am J Surg Pathol, 2012, 36(7):993-999. http://cn.bing.com/academic/profile?id=fd7c6abb007f0ea4ccaf0809a9f4fd89&encoded=0&v=paper_preview&mkt=zh-cn [7] Russell-Goldman E, Hornick JL, Qian X, et al. NKX2.2 immunohistochemistry in the distinction of Ewing sarcoma from cytomorphologic mimics:Diagnostic utility and pitfalls[J]. Cancer Cytopathol, 2018, 126(11):942-949. http://www.researchgate.net/publication/328616987_NKX22_immunohistochemistry_in_the_distinction_of_Ewing_sarcoma_from_cytomorphologic_mimics_Diagnostic_utility_and_pitfalls [8] Bledsoe KL, McGee-Lawrence ME, Camilleri ET, et al. RUNX3 facilitates growth of Ewing sarcoma cells[J]. J Cell Physiol, 2014, 229(12):2049-2056. http://smartsearch.nstl.gov.cn/paper_detail.html?id=bba955814a7eca23574e0f6b35337355 [9] Lessnick SL, Ladanyi M. Molecular pathogenesis of Ewing sarcoma:new therapeutic and transcriptional targets[J]. Annu Rev Pathol, 2012, 7:145-159. http://europepmc.org/articles/PMC3555146/ [10] Renzi S, Anderson ND, Light N, et al. Ewing-like sarcoma:An emerging family of round cell sarcomas[J]. J Cell Physiol, 2019, 234(6):7999-8007. http://www.onacademic.com/detail/journal_1000040857199410_50b7.html [11] Chen S, Deniz K, Sung YS, et al. Ewing sarcoma with ERG gene rearrangements:A molecular study focusing on the prevalence of FUS-ERG and common pitfalls in detecting EWSR1-ERG fusions by FISH[J]. Genes Chromosomes Cancer, 2016, 55(4):340-349. http://cn.bing.com/academic/profile?id=fc13a95b1f3cf0424b74d9ddc80b5056&encoded=0&v=paper_preview&mkt=zh-cn [12] Brohl AS, Solomon DA, Chang W, et al. The genomic landscape of the Ewing sarcoma family of tumors reveals recurrent STAG2 mutation[J]. PLoS Genet, 2014, 10(7):e1004475. http://www.wanfangdata.com.cn/details/detail.do?_type=perio&id=Doaj000004799187 [13] Lindén M, Thomsen C, Grundevik P, et al. FET family fusion oncoproteins target the SWI/SNF chromatin remodeling complex[J]. EMBO Rep, 2019, 20(5):e45766. http://www.ncbi.nlm.nih.gov/pubmed/30962207 [14] Casali PG, Bielack S, Abecassis N, et al. Bone sarcomas:ESMO-paedCan-EURACAN clinical practice guidelines for diagnosis, treatment and follow-up[J]. Ann Oncol, 2018, 29(Suppl 4):iv79-iv95. http://www.researchgate.net/publication/328104350_Bone_sarcomas_ESMO-PaedCan-EURACAN_Clinical_Practice_Guidelines_for_diagnosis_treatment_and_follow-up [15] Theisen ER, Pishas KI, Saund RS, et al. Therapeutic opportunities in Ewing sarcoma:EWS-FLI inhibition via LSD1 targeting[J]. Oncotarget, 2016, 7(14):17616-17630. http://pubmedcentralcanada.ca/pmcc/articles/PMC4951237/ [16] Mastrangelo T, Modena P, Tornielli S, et al. A novel zinc finger gene is fused to EWS in small round cell tumor[J]. Oncogene, 2000, 19(33):3799-3804. http://www.wanfangdata.com.cn/details/detail.do?_type=perio&id=79ac41e33942e10329068cb3895092c1 [17] Yamaguchi S, Yamazaki Y, Ishikawa Y, et al. EWSR1 is fused to POU5F1 in a bone tumor with translocation t(6;22)(p21;q12)[J]. Genes Chromosomes Cancer, 2005, 43(2):217-222. http://europepmc.org/abstract/MED/15729702 [18] Wang L, Bhargava R, Zheng T, et al. Undifferentiated small round cell sarcomas with rare EWS gene fusions:identification of a novel EWSSP3 fusion and of additional cases with the EWS-ETV1 and EWS-FEV fusions[J]. J Mol Diagn, 2007, 9(4):498-509. http://www.sciencedirect.com/science/article/pii/S1525157810604229 [19] Szuhai K, Ijszenga M, de Jong D, et al. The NFATc2 gene is involved in a novel cloned translocation in a Ewing sarcoma variant that couples its function in immunology to oncology[J]. Clin Cancer Res, 2009, 15(7):2259-2268. http://www.wanfangdata.com.cn/details/detail.do?_type=perio&id=d7109dc60577c8e8bd1ae971bb70ec30 [20] Koelsche C, Kriegsmann M, Kommoss F, et al. DNA methylation profiling distinguishes Ewing-like sarcoma with EWSR1-NFATc2 fusion from Ewing sarcoma[J]. J Cancer Res Clin Oncol, 2019, 145(5):1273-1281. http://www.ncbi.nlm.nih.gov/pubmed/30895378 [21] Bode-Lesniewska B, Fritz C, Exner GU, et al. EWSR1-NFATC2 and FUSNFATC2 gene fusion-associated mesenchymal tumors:clinicopathologic correlation and literature review[J]. Sarcoma, 2019, 2019:9386390. https://www.ncbi.nlm.nih.gov/pubmed/31049020 [22] Charville GW, Wang WL, Ingram DR, et al. PAX7 expression in sarcomas bearing the EWSR1-NFATC2 translocation[J]. Mod Pathol, 2019, 32(1):154-156. http://www.wanfangdata.com.cn/details/detail.do?_type=perio&id=2ab9a3597357c3740a065800e19658e4 [23] Yoshida KI, Machado I, Motoi T, et al. NKX3-1 is a useful immunohistochemical marker of EWSR1-NFATC2 sarcoma and mesenchymal chondrosarcoma[J]. Am J Surg Pathol, 2020, 44(6):719-728. http://www.wanfangdata.com.cn/details/detail.do?_type=perio&id=10.1097/PAS.0000000000001441 [24] Sumegi J, Nishio J, Nelson M, et al. A novel t(4;22)(q31;q12) produces an EWSR1-SMARCA5 fusion in extraskeletal Ewing sarcoma/primitive neuroectodermal tumor[J]. Mod Pathol, 2011, 24(3):333-342. http://www.wanfangdata.com.cn/details/detail.do?_type=perio&id=607d4df9a23bab604240bc2b5e808260 [25] Bridge JA, Sumegi J, Druta M, et al. Clinical, pathological, and genomic features of EWSR1-PATZ1 fusion sarcoma[J]. Mod Pathol, 2019, 32(11):1593-1604. http://www.nature.com/articles/s41379-019-0301-1 [26] Min L, Shen J, Tu C, et al. The roles and implications of exosomes in sarcoma[J]. Cancer Metastasis Rev, 2016, 35(3):377-390. http://www.wanfangdata.com.cn/details/detail.do?_type=perio&id=7a760bbd06f83faa9c5dee0ff201ea8f [27] Richkind KE, Romansky SG, Finklestein JZ. t(4;19)(q35;q13.1):a recurrent change in primitive mesenchymal tumors[J]? Cancer Genet Cytogenet, 1996, 87(1):71-74. http://www.sciencedirect.com/science/article/pii/0165460896853222 [28] Haidar A, Arekapudi S, DeMattia F, et al. High-grade undifferentiated small round cell sarcoma with t(4;19)(q35;q13.1) CIC-DUX4 fusion:emerging entities of soft tissue tumors with unique histopathologic features-a case report and literature review[J]. Am J Case Rep, 2015, 16:87-94. http://www.ncbi.nlm.nih.gov/pubmed/25683183 [29] Italiano A, Sung YS, Zhang L, et al. High prevalence of CIC fusion with double-homeobox (DUX4) transcription factors in EWSR1-negative undifferentiated small blue round cell sarcomas[J]. Genes Chromosomes Cancer, 2012, 51(3):207-218. http://www.wanfangdata.com.cn/details/detail.do?_type=perio&id=4c72bcdaef5147add911fed2c8274263 [30] Le Loarer F, Pissaloux D, Watson S, et al. Clinicopathologic features of CIC-NUTM1 sarcomas, a new molecular variant of the family of CICfused sarcomas[J]. Am J Surg Pathol, 2019, 43(2):268-276. http://journals.lww.com/ajsp/Abstract/2019/02000/Clinicopathologic_Features_of_CIC_NUTM1_Sarcomas,.15.aspx [31] Antonescu CR, Owosho AA, Zhang L, et al. Sarcomas with CIC-rearrangements are a distinct pathologic entity with aggressive outcome:a clinicopathologic and molecular study of 115 Cases[J]. Am J Surg Pathol, 2017, 41(7):941-949. http://cn.bing.com/academic/profile?id=faecc198382b912f54b7313ce0bc255d&encoded=0&v=paper_preview&mkt=zh-cn [32] Siegele B, Roberts J, Black JO, et al. DUX4 immunohistochemistry is a highly sensitive and specific marker for CIC-DUX4 fusion-positive round cell tumor[J]. Am J Surg Pathol, 2017, 41(3):423-429. http://cn.bing.com/academic/profile?id=b1f998f9eeadb05a5d32dfc9b8b1c3ff&encoded=0&v=paper_preview&mkt=zh-cn [33] Le Guellec S, Velasco V, Pérot G, et al. ETV4 is a useful marker for the diagnosis of CIC-rearranged undifferentiated round-cell sarcomas:a study of 127 cases including mimicking lesions[J]. Mod Pathol, 2016, 29(12):1523-1531. http://www.nature.com/modpathol/journal/v29/n12/abs/modpathol2016155a.html [34] Yoshimoto T, Tanaka M, Homme M, et al. CIC-DUX4 induces small round cell sarcomas distinct fromEwingsarcoma[J]. Cancer Res, 2017, 77(11):2927-2937. http://www.ncbi.nlm.nih.gov/pubmed/28404587 [35] Pierron G, Tirode F, Lucchesi C, et al. A new subtype of bone sarcoma defined by BCOR-CCNB3 gene fusion[J]. Nat Genet, 2012, 44(4):461-466. http://www.ncbi.nlm.nih.gov/pubmed/22387997 [36] Le Loarer F, Pissaloux D, Coindre JM, et al. Update on families of round cell sarcomas other than classical Ewing sarcomas[J]. Surg Pathol Clin, 2017, 10(3):587-620. http://www.wanfangdata.com.cn/details/detail.do?_type=perio&id=97039542443172f9421e70d3fa024fd8 [37] Specht K, Zhang L, Sung YS, et al. Novel BCOR-MAML3 and ZC3H7BBCOR gene fusions in undifferentiated small blue round cell sarcomas[J]. Am J Surg Pathol, 2016, 40(4):433-442. http://smartsearch.nstl.gov.cn/paper_detail.html?id=7ff1b03f7ee3b4e7a02d9184b80d78a8 [38] Kao YC, Owosho AA, Sung YS, et al. BCOR-CCNB3 fusion positive sarcomas:a clinicopathologic and molecular analysis of 36 cases with comparison to morphologic spectrum and clinical behavior of other round cell sarcomas[J]. Am J Surg Pathol, 2018, 42(5):604-615. http://smartsearch.nstl.gov.cn/paper_detail.html?id=eb70811bf9de7a2bb7b42a9657bbc378 [39] Matsuyama A, Shiba E, Umekita Y, et al. Clinicopathologic diversity of undifferentiated sarcoma with BCOR-CCNB3 fusion:analysis of 11 cases with a reappraisal of the utility of immunohistochemistry for BCOR and CCNB3[J]. Am J Surg Pathol, 2017, 41(12):1713-1721. http://www.ncbi.nlm.nih.gov/pubmed/28877060 [40] Kao YC, Sung YS, Zhang L, et al. BCOR overexpression is a highly sensitive marker in round cell sarcomas with BCOR genetic abnormalities[J]. Am J Surg Pathol, 2016, 40(12):1670-1678. http://cn.bing.com/academic/profile?id=6e9cd444d49de4540f7e050040930ac4&encoded=0&v=paper_preview&mkt=zh-cn [41] Puls F, Niblett A, Marland G, et al. BCOR-CCNB3(Ewing-like) sarcoma:a clinicopathologic analysis of 10 cases, in comparison with conventional Ewing sarcoma[J]. Am J Surg Pathol, 2014, 38(10):1307-1318. http://europepmc.org/abstract/med/24805859 [42] LudwigK, AlaggioR, ZinA, et al. BCOR-CCNB3undifferentiatedsarcomadoes immunohistochemistry help in the identification[J]? Pediatr Dev Pathol, 2017, 20(4):321-329. http://www.onacademic.com/detail/journal_1000040039441810_5f3e.html -
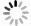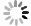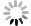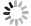



 下载:
下载: